
To assess these distance determination methods, hypothetical experimental data, including random noise and peak overlap, are calculated for an arbitrary true protein structure. The value more » of this additional complexity is also addressed. Iterative relaxation matrix calculations, which account for dipolar interactions between all protons in a molecule, can accurately determine internuclear distances with little or no a priori knowledge of the molecular structure. Effects of these systematic errors on the resulting protein structure are examined. Although the simple isolated spin pair approximation (ISPA) generally used can result in systematic errors in distances, the large number of constraints enables proteins structure to be defined with reasonably high resolution. Solution structures for many proteins have been determined to date utilizing interproton distance constraints estimated from two-dimensional nuclear Overhauser effect (2D NOE) spectra.
Eventhough the method requires one matrix inverse for each time interval oftomographic acquisition, efficient estimates of the tissue kineticparameters in a dynamic cardiac SPECT study can be obtained with presentday desk-top computers. Numerical methods are used to calculate thesecond derivative of the chi-square criterion to obtain estimates of thecovariance matrix for the weighted least square parameter estimates. A one-compartment model is fit to the tissueactivity curves assuming a noisy blood input function to give weightedleast squares estimates of blood volume fraction, wash-in and wash-outrate constants specifying the kinetics of 99mTc-teboroxime for theleftventricular myocardium. Also, curves for the variance of thetwo estimates of the sum and for the covariance between the two ROIestimates are generated as a function of time at convergence using anexpression obtained from the fixed-point solution of the statisticalerror of the reconstruction. Time-activity curves for a sumof activity in a blood region inside the left ventricle and a sum in acardiac tissue region are generated. For each transaxial slicesets of sequential tomographic projections are reconstructed into asequence of transaxial reconstructions more » usingfor each reconstruction inthe time sequence an iterative MAP reconstruction to calculate themaximum a priori reconstructed estimate. This paper provides a method for calculating thecovariance matrix of the kinetic parameters, which are determined usingweighted least squares fitting that incorporates the estimated varianceand covariance of the dynamic reconstructions. In dynamic cardiac SPECT estimates of kinetic parameters ofa one-compartment perfusion model are usually obtained in a two stepprocess: 1) first a MAP iterative algorithm, which properly models thePoisson statistics and the physics of the data acquisition, reconstructsa sequence of dynamic reconstructions, 2) then kinetic parameters areestimated from time activity curves generated from the dynamicreconstructions. In order to prove these results, we introduce a new class of events which we call decoupling events and two inequalities for these events.
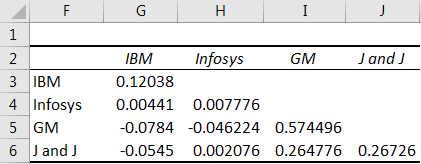
Covariance matrix free#
We also introduce DLR equations for the random cluster model and use them to establish ergodicity of the free measure.
